valuesets
A LinkML-based Framework for Standardized Enumerations with Semantic Grounding
Christopher J. Mungall Lawrence Berkeley National Laboratory
The Problem: Data Standardization is Hard
Every project reinvents the wheel with inconsistent representations:
# Different datasets, same concept
vital_status = "alive" # Dataset A
vital_status = "LIVING" # Dataset B
vital_status = 1 # Dataset C
vital_status = "A" # Dataset D
Result: Thousands of incompatible representations blocking data integration
The Semantic Chasm
Despite massive infrastructure investment: - NLM VSAC: 1,520+ clinical value sets - NCI Thesaurus: 192,000 cancer concepts - NIH CDEs: 142,000+ common data elements - HL7 FHIR: Healthcare terminology standards
Yet: Scientific software still uses ad-hoc enumerations
Why? Complexity gap between terminology services and everyday programming
The Gap: What Exists vs What Developers Need
| Existing Systems Provide | Developers Actually Need |
|---|---|
| Runtime services | Compile-time artifacts |
| Comprehensive coverage | Common values quickly |
| Authentication & servers | Zero dependencies |
| Healthcare-focused | Cross-domain support |
| Complex APIs | Native enums with IDE support |
valuesets: Bridging the Gap
Core Idea: Compile semantically-grounded value sets into type-safe native code
A collection of common, standardized enumerations that: - Link every value to ontology terms - Provide Python-first convenience with multi-language support - Built on LinkML standards - Have zero runtime dependencies
"Stealth Semantics" in Action
from valuesets.enums.core import VitalStatusEnum
status = VitalStatusEnum.ALIVE
print(status.value) # "ALIVE"
print(status.get_meaning()) # "NCIT:C37987"
print(status.get_description()) # "Living or alive"
# Semantic interoperability across systems
if status1.get_meaning() == status2.get_meaning():
process_compatible_records()
Simple interface, semantic power when needed
Rich Metadata & Ontology Mappings
from valuesets.enums.bio.structural_biology import StructuralBiologyTechnique
technique = StructuralBiologyTechnique.CRYO_EM
print(technique.get_description())
# "Cryo-electron microscopy"
print(technique.get_meaning())
# "CHMO:0002413" (Chemical Methods Ontology)
print(technique.get_annotations())
# {'resolution_range': '2-30 Å typical', ...}
Cross-Domain Coverage
322 enumerations across 22 domains:
- Biology (127): Taxonomy, cell biology, structural techniques
- Physical Sciences (48): Chemical elements, materials, structures
- Data Science (43): Statistical tests, ML models, quality metrics
- Healthcare (29): Clinical findings, vital status, demographics
- Computing (23): File formats, languages, maturity levels
- Geographic & Temporal (31): Countries, time zones, spatial relations
78% have ontology mappings → 8,743 semantic links
Architecture: Build-Time not Runtime
┌─────────────────┐
│ LinkML YAML │ ← Human-editable schemas
│ (source) │
└────────┬────────┘
│
▼
┌─────────────────┐
│ Code Generators │ ← Transform to multiple formats
└────────┬────────┘
│
├──→ Python (Pydantic enums)
├──→ TypeScript (type-safe enums)
├──→ JSON Schema (validation)
├──→ OWL (semantic web)
└──→ SQL DDL (database constraints)
Progressive Semantic Enhancement
Three levels of usage:
-
~90% of use cases: Simple type-safe enumerations
python status = VitalStatusEnum.ALIVE # Just works -
~9% of use cases: Access metadata for UIs/docs
python label = status.get_description() -
~1% of use cases: Full semantic integration
python ontology_term = status.get_meaning() # "NCIT:C37987"
Comparison with Established Systems
| System | Scope | Access | Size | Advantage |
|---|---|---|---|---|
| VSAC | Clinical QM | API (auth) | Service | Cross-domain, no auth |
| NCIt | Cancer | 500MB OWL | Service | Lightweight, simple |
| FHIR | Healthcare | Term. server | Service | Compile-time, no server |
| valuesets | Cross-domain | Native packages | <50MB | Developer-friendly |
valuesets complements not replaces - provides practical bridge
Design Principles
- Semantic Grounding: Every value links to ontologies
- Developer Ergonomics: Native enums, full IDE support
- Modular Organization: Import only what you need
- Extensibility: Add new enums without breaking changes
- Multi-format: JSON Schema, OWL, SQL, native code
- FAIR Compliance: Persistent IDs, metadata, open access
LinkML: The Foundation
enums:
VitalStatusEnum:
description: Status indicating whether individual is alive or deceased
permissible_values:
ALIVE:
description: Living or alive
meaning: NCIT:C37987
DECEASED:
description: Dead or deceased
meaning: NCIT:C28554
UNKNOWN:
description: Vital status is not known
meaning: NCIT:C17998
Human-readable → Machine-processable → Multiple outputs
Integration Patterns
Four primary adoption strategies:
- Direct Adoption: Greenfield projects
- Mapping Layer: Legacy system translation
- Hybrid Approach: Dev/test vs. production
- Semantic Bridge: Ontology integration
All paths support incremental adoption
FAIR Data Principles
valuesets is FAIR-compliant:
- Findable: w3id.org permalinks, rich metadata
- Accessible: Open source, multiple formats
- Interoperable: LinkML, OWL, JSON-LD, FHIR
- Reusable: Clear licensing, documented provenance
Published as OWL ontology: https://w3id.org/valuesets/valuesets.owl.ttl
OWL Rendering in Protege
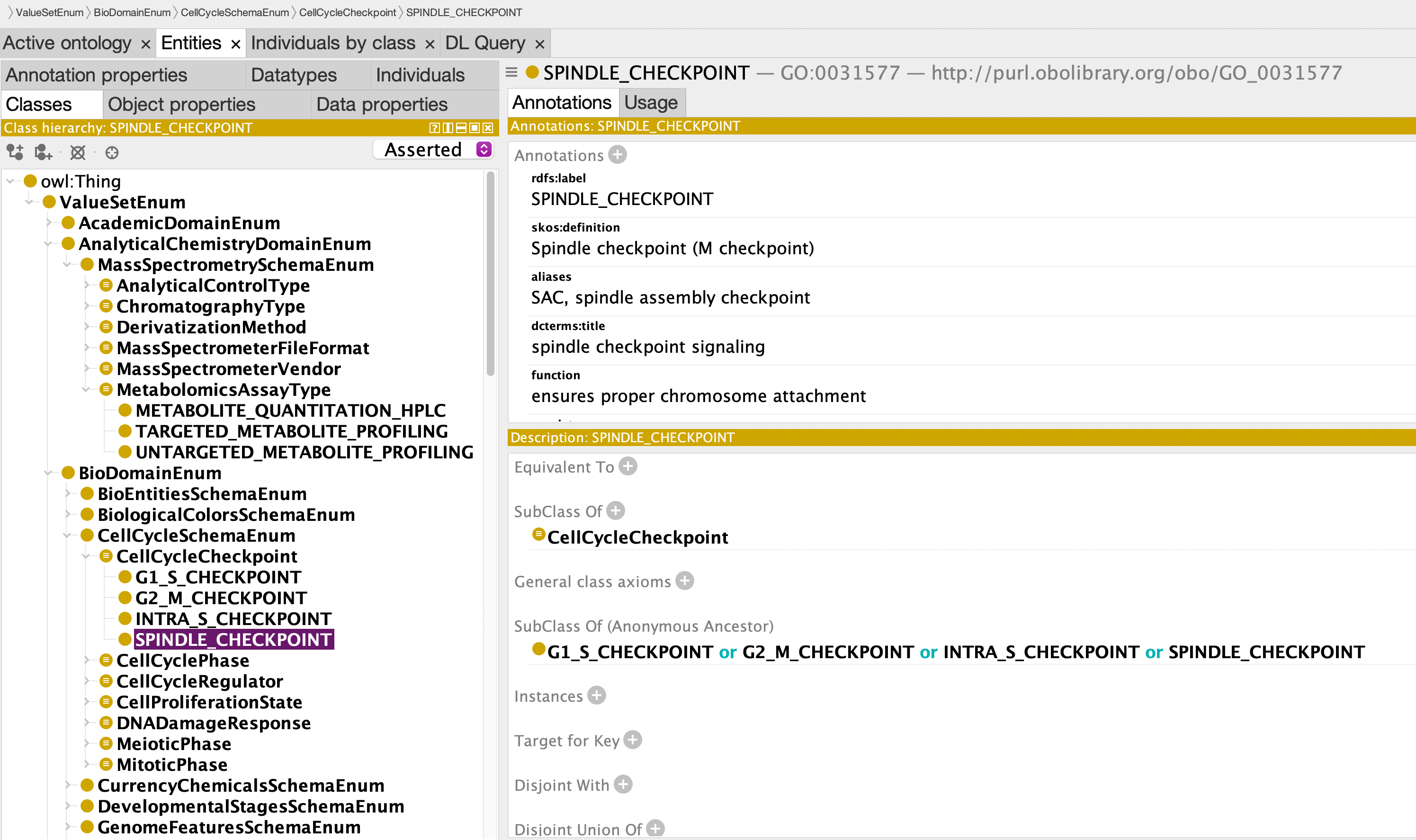
Value sets as OWL classes with rich semantic annotations: - Hierarchical organization (e.g., CellCycleCheckpoint > SPINDLE_CHECKPOINT) - Ontology mappings (GO:0031577) - Definitions, aliases, and functional descriptions - Browsable in standard ontology tools (Protege, OLS, BioPortal)
Quality Assurance
Automated validation on every commit:
| Validation Type | Coverage | Purpose |
|---|---|---|
| Syntax | 100% schemas | LinkML compliance |
| Semantic | All mappings | Ontology term verification |
| Cross-reference | All namespaces | External reference resolution |
| Completeness | All enums | Missing descriptions/mappings |
| Consistency | All values | Duplicate detection |
Dynamic Enums (Coming Soon)
Current: Static values with ontology mappings
Future: Runtime expansion from ontologies
CellTypeEnum:
reachable_from:
source_ontology: obo:cl
source_nodes:
- CL:0000540 # neuron
relationship_types:
- rdfs:subClassOf
Hybrid: static core + dynamic expansion for comprehensive coverage
Future Directions
Coverage Expansion: - Social sciences, engineering, humanities
Technical Enhancements: - Web-based validation APIs - AI-assisted mapping tools - Enhanced dynamic enum support
Governance: - Domain-specific editorial committees - Formal deprecation policies - Maturity level indicators
Sustainability Through Simplicity
Technical Sustainability: - Zero runtime dependencies - No infrastructure costs - Works offline, on laptops
Social Sustainability: - Low barrier to contribution (edit YAML) - Community-driven development - Clear exit strategies
Economic Sustainability: - No hosting/licensing costs - Open source model
By the Numbers
| Metric | Count |
|---|---|
| Enumerations | 322 |
| Permissible Values | 7,512 |
| Ontology Namespaces | 117 |
| Semantic Mappings | 8,743 |
| Domain Modules | 22 |
| Schemas | 68 |
| Ontology Mapping Coverage | 78% |
Quick Start
# Install
pip install valuesets
# Use immediately
from valuesets.enums.bio.taxonomy import CommonOrganismTaxaEnum
from valuesets.enums.core import VitalStatusEnum
human = CommonOrganismTaxaEnum.HUMAN
print(human.get_meaning()) # "NCBITaxon:9606"
status = VitalStatusEnum.ALIVE
print(status.get_meaning()) # "NCIT:C37987"
5-minute experience: Value within 5 minutes of discovery
Contributing
We welcome contributions from:
- Domain Experts: Add value sets for your field
- Developers: Improve tooling, fix issues
- Users: Report missing enums, share use cases
Process: 1. Edit YAML schema files 2. Add ontology mappings (use OLS) 3. Include descriptions and examples 4. Submit pull request
See: CONTRIBUTING.md
Development Commands
# Using just command runner
just --list # Show all commands
just test # Run tests
just doctest # Run doctests
just validate # Validate schemas
just site # Build documentation
All managed through modern development workflows
Resources
- Docs: https://linkml.io/valuesets/
- Repository: https://github.com/linkml/common-value-sets
- PyPI: https://pypi.org/project/valuesets/
- OWL Ontology: https://w3id.org/valuesets/valuesets.owl.ttl
- LinkML: https://linkml.io/
Key Insight
"The problem is not missing standards but mismatched abstractions"
valuesets bridges the gap: - Terminology services → Terminology artifacts - Runtime flexibility → Compile-time guarantees - Institutional deployment → Developer laptops
Making semantic standards accessible to everyday programming
valuesets
Making Data Standardization Simple, Semantic, and Scalable
Try it today:
pip install valuesets
Questions? Christopher J. Mungall • cjmungall@lbl.gov Lawrence Berkeley National Laboratory
Appendix: Example Domains
Biological Sciences: - Taxonomy (NCBI), Cell types (CL), Cell cycle (GO) - Gene Ontology evidence codes - Structural biology techniques (CHMO) - Model organisms
Data Science: - Statistical tests (STATO) - ML model types, dataset splits - Data quality indicators
Clinical/Healthcare: - Vital status (NCIT), blood types (SNOMED) - Marital status, employment status - Healthcare encounter types
Appendix: Semantic Web Integration
from valuesets.enums.bio.cell_cycle import CellCyclePhase
# Generate SPARQL query
phase = CellCyclePhase.S_PHASE
go_term = phase.get_meaning() # "GO:0000084"
sparql = f"""
SELECT ?gene ?function
WHERE {{
?gene cellCyclePhase <{go_term}> .
?gene hasFunction ?function .
}}
"""
Seamless integration with knowledge graphs
Appendix: Multi-Language Support
Current: - Python (Pydantic enums) - TypeScript (type-safe) - JSON Schema - OWL/RDF
Planned: - Java - R - Julia - Rust
LinkML generates idiomatic code for each language
Credits & Acknowledgments
Contributors: - Christopher J. Mungall - Lawrence Berkeley National Laboratory - Justin Reese - Lawrence Berkeley National Laboratory
Built with: - LinkML - Linked Data Modeling Language - linkml-project-copier - Project template - OBO Foundry - Biological ontologies - OLS/BioPortal - Ontology lookup services
Open source • MIT License • Community-driven